Could you give us some background information concerning your project, and why it is important?
The discovery of antibiotics has revolutionized the treatment of bacterial infections in human and veterinary medicine; the world’s major anti-infective agents are β-lactam antibiotics.
Cephalosporin and penicillin derivatives play a major role in the treatment of bacterial infection and are produced in industrial scale by fermentation of the two taxonomically non-related fungi Acremonium chrysogenum and Penicillium chrysogenum. Both were subjects of long-term strain development programs to reach economically relevant antibiotic titres.
We aim to identify factors that have an impact on the biosynthesis of β-lactam antibiotics in both organisms, in order to develop strategies for further strain improvement.
These programs resulted in alterations that are visible at the genome, transcriptome and proteome level of industrial strains. The molecular details of these changes are widely unknown, but might provide concepts to improve antibiotic titres of production strains by targeted genetic engineering. We aim to identify factors that have an impact on the biosynthesis of β-lactam antibiotics in both organisms, in order to develop strategies for further strain improvement.
Could you briefly describe your research and your findings?
It was so far unknown, whether equivalent changes in gene expression lead to elevated titres in production strains from both fungi. To address this question we have performed a comparative gene expression analysis of wild-type and derived production strains.
For both fungi, we showed that the expression levels of a wide range of genes were affected by strain improvement programs. Further, we observed an upregulation of genes belonging to primary and secondary metabolism, notably those involved in precursor supply for β-lactam production; in contrast, genes not essential for β-lactam production were downregulated.
Examples for downregulated genes are those involved in stress response or transport processes which are dispensable under the controlled conditions of industrial fermentations. In addition to that we investigated deletion mutants from both production strains, lacking Velvet a major global transcriptional regulator.
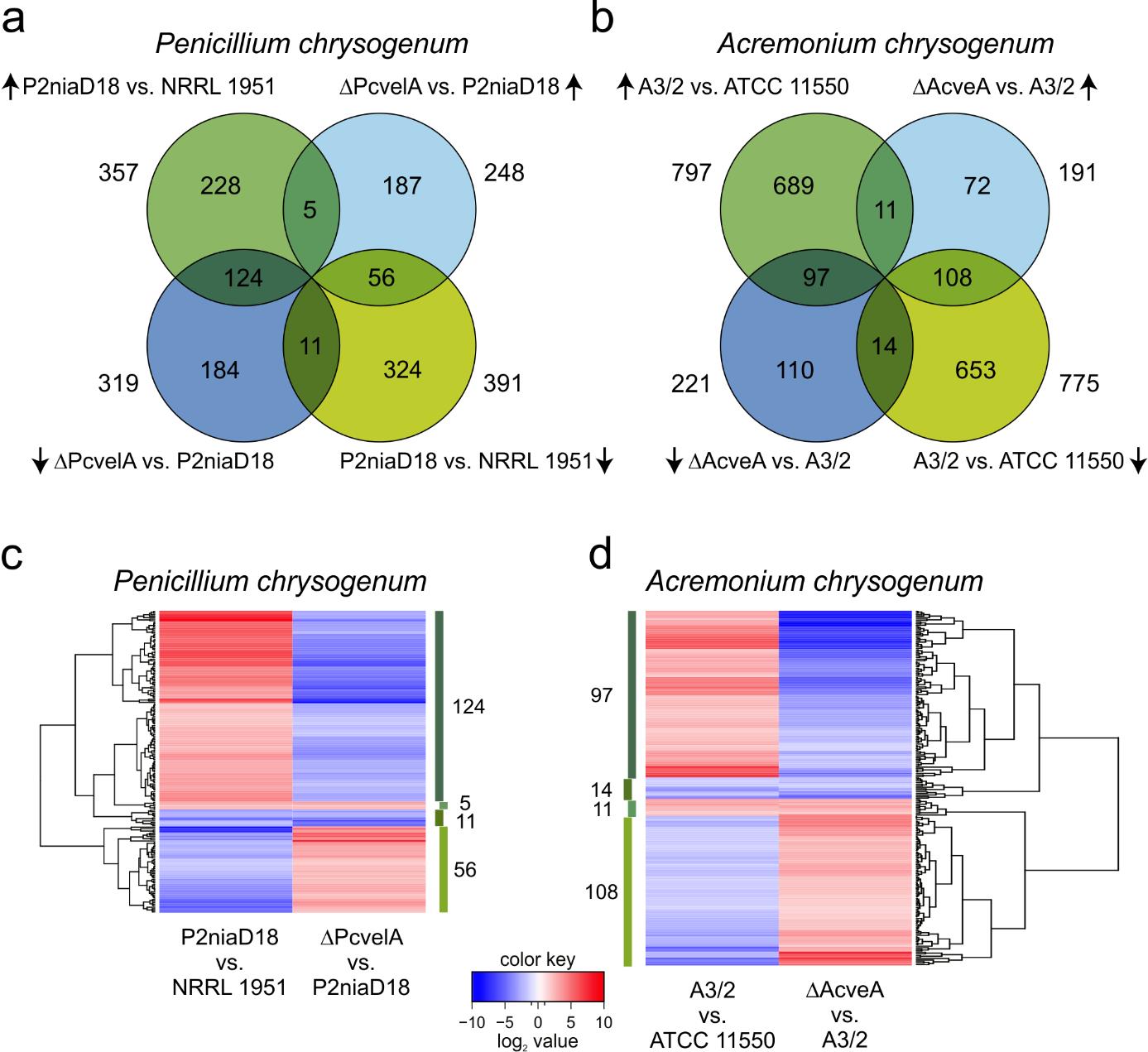
We showed that Velvet regulates nearly 50% of all secondary metabolite clusters in both organisms and most importantly, 35-57% of Velvet target genes are among those that showed differential expression in both improved industrial strains. Thus, the major finding of our comparative transcriptome analysis is that strain improvement programs in two unrelated fungal β-lactam antibiotic producers alter the expression of target genes of Velvet, a global regulator of secondary metabolism.
From these results, we conclude that regulatory alterations are crucial contributing factors for improved β-lactam antibiotic titres during strain improvement.
What brought you to work with fungi? And what do you enjoy most about your research?
I really like to work in applied research, knowing that results might have a direct impact on applied microbiology.
I started to work with fungal β-lactam antibiotic producers during my bachelor thesis in the group of Prof. Dr. Ulrich Kück at Bochum University. I was always fascinated by the regulatory networks of filamentous fungi and how they managed to integrate biosynthetic pathways of prokaryotic origin within their metabolism. Furthermore, I really like to work in applied research, knowing that results might have a direct impact on applied microbiology.
Where would you like to see your research taking you in the next few years?
With the data sets obtained from A. chrysogenum and P. chrysogenum, we have identified highly interesting target genes for further analysis and hopefully are able to resolve their impact on β-lactam antibiotic biosynthesis to obtain a deeper understanding of this important process.
What advice would you give young scientists starting out in research?
I learnt during my PhD many novel aspects of microbiology and know now that studying biology was the right choice for me. I really love the combination of applied and theoretical biology and the option to constantly discover something new for my PhD thesis. Therefore, I encourage everyone interested in biology to start their study.
Enjoyed reading this blog? Read our other Fungal Genetics Conference Q&As here:
Comments