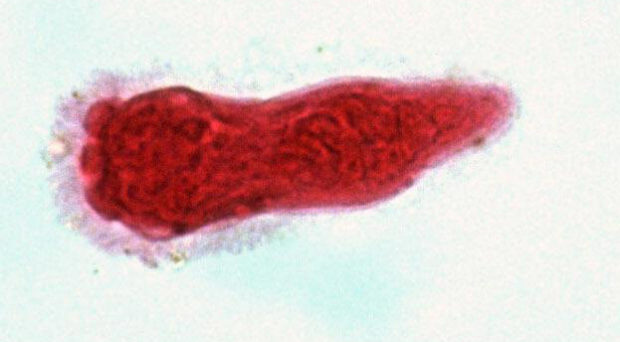
Parasitic worms belong to one of two helminth phyla – Nematoda and Platyhelminthes. Nematodes are incredibly diverse and are found nearly everywhere – from cold seeps in the deep ocean to the Dry Valleys of Antarctica, and have been found three kilometres deep in the Earth’s crust. Nematodes were even found alive in the debris of Space Shuttle Columbia after its tragic breakup re-entering the earth’s atmosphere in 2003. Although some nematodes are free-living, many are parasites of animals, plants and fungi. Phylum Platyhelminthes contains at least 15,000 species of known parasites, grouped into two parasitic clades: tapeworms (Cestoda) and flukes (Trematoda). Platyhelminthes also includes the well-known (non-parasitic) flatworms (Planaria).
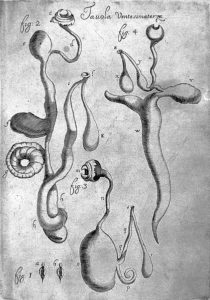
The impact of these worms on humans is profound: parasitic helminths infect up to a quarter of the earth’s population and cause diseases that are often disfiguring, debilitating, and painful. These include schistosomiasis, elephantiasis, river blindness, and many others that often disproportionately impact low-income countries. Furthermore, helminths are formidable agricultural pathogens, causing production losses in livestock and crop farming.
In a recent paper in Nature Genetics, a group of scientists sequenced, assembled and annotated the genomes of 45 helminth species, more than doubling the number of available helminth genomes. The goal was to compare genes and gene families between species to help identify common features, potential drug targets, and insights into the processes involved in parasitism.
Using 81 helminth genomes (including genomes previously published), a total of 1.6 million genes were collected, of which 1.4 million were placed into ~100,000 gene families (genes that are related through sharing a common ancestor). The evolutionary history of these genes was studied by comparing them between different species. A key aspect of this was looking for expansions that have taken place within gene families. Imagine you were trying to identify carpenters by comparing carpenters to laypeople. If you looked in my toolbox, you would find some dust, an ancient hammer and a screwdriver. However, if you were to look at a carpenter’s toolbox, you would see a large number of specialised and diverse (yet related) tools. Using this logic, you could conceivably identify potential carpenters by looking at the functional characteristics of carpenters relative to unskilled laypeople.
In the same way that the carpenter’s repertoire is far larger relative to the meagre complement of tools in my cupboard, identification of gene expansions in a group of species relative to another indicates that this group of genes is doing something useful (or at least interesting) and may warrant further investigation. Not that single genes don’t do important things – expansions are just easier to spot.
Parasitism has evolved independently in multiple helminth lineages. The evolution of genes and metabolic pathways in these species appears to have reflected this, with the authors reporting expansion in gene families that are involved in parasitic processes, as well as the potential alteration of parasite metabolism to suit the worm’s environment. Interestingly the authors found instances of gene expansion and metabolic alteration that have occurred independently, but share characteristics. Such instances suggest that, although parasitism has evolved multiple times in different species, common features of their parasitic lifestyle have driven the development of adaptations that resemble each other. For example, if two species have evolved (independently) to be parasitic, they will both need to avoid the host’s immune response. As the two different species have to do similar things, the adaptations may resemble each other.
An example of this is the expansion of the SCP/TAPS gene family in Nematodes. This gene family, which has been widely implicated in helminth parasite pathogenesis, has undergone large-scale expansions in multiple independent parasitic nematode groups. Another example is the independent loss of a glycine cleavage pathway in Trichinella and the filaria, both of which are tissue-dwelling parasites.
The authors also wanted to use the genomic dataset to identify potential anthelmintic drugs and drug targets. To this effect, the authors screened ChEMBL – a database of medicinal and bio-active compounds, with helminth protein sequences, to see whether they matched any of the potential drug targets in the database. The final result set included a library of 5082 drugs and drug-like compound. After the results were refined, they identified 40 high-priority drug targets associated with 720 drugs and drug-like compounds. These data serve to inform the repurposing of existing drugs and drugs undergoing trial, as well as providing a broader base of medicinal compounds for anthelmintic screening.
The increasing ease and decreasing cost of whole genome sequencing has led to an embarrassment of riches with regards to biological sequence data. Here, the authors present an example of how these data can be so valuable, using a considerable body of helminth genome sequence to provide insights into the genetic and metabolic basis of helminth parasitism, as well as identifying a battery of potential therapeutic compounds. Studies like this will increase in number and power as sequence data continues to accumulate, allowing finer comparisons between more closely-related taxa.

Comments